2021.06.01.4
Files > Volume 6 > Vol 6 No 1 2021
INVESTIGATION / RESEARCH
Genetic transformation of apical meristematic shoots in the banana cultivar ‘Williams’
Liliana Villao1, José Flores1, 2, Efrén Santos-Ordóñez 1, 3
Available from: http://dx.doi.org/10.21931/RB/2021.06.01.4
ABSTRACT
Bananas and plantains (Musa spp.) are among the most critical socioeconomic crops globally, being a staple food for millions of people in the tropics and an essential component for the export market, including the subtropics. Besides conventional breeding, genetic improvement of bananas and plantains could be performed through genetic engineering and new breeding techniques. Furthermore, plant tissue culture is essential for these technologies, including developing embryogenic cell suspensions and in vitro plants. The transient and stable genetic transformation could be performed from in vitro plants, shortening Musa transgenic lines development compared to genetic transformation while using embryogenic cell suspension. In this study, a genetic transformation protocol was established from banana apical meristems for the ‘Williams’ cultivar (genotype AAA). The protocol was based on the co-cultivation of the explants (whole in vitro plants or bisected meristematic tissues derived from in vitro plants) with Agrobacterium tumefaciens strain LBA4404 harboring two binary vectors denominated pLVCIBE1 (cassette: MabHIPP promoter::luc2::Tnos, P35S::hpt::Tnos) and pLVCIBE2 (cassette: P35S::luc2::Tnos, P35S::hpt::Tnos), independently. The stable genetic transformation was obtained by subculturing in vitro banana plants in selection medium (12.5µg/mL of hygromycin) for 8 weeks from bisected meristematic tissue transformation. Genetic transformation was confirmed in vivo with the use of the luciferase reporter gene system. Furthermore, PCR was performed on DNA extracted from leaves of regenerated transgenic in vitro plants after 8 weeks of selection, confirming stable genetic transformation. Therefore, genetic transformation was achieved in the apical meristematic tissue of in vitro banana plants with co-cultivation of Agrobacterium tumefaciens.
Keywords: Agrobacterium tumefaciens, luciferase, micropropagation, promoter
INTRODUCTION
Banana and plantain crops (Musa spp.) have great importance in the tropics and some subtropical regions worldwide due to the export market and food security. In Ecuador, this fruit's consumption is massive and form part of the daily diet for millions of people 1. Banana and plantain crops are affected by biotic and abiotic stresses. Genetic improvement arises as an essential strategy to overcome reduced productivity due to different stresses in the crops. Musa's conventional breeding is difficult due to different factors including high level of sterility, interploidy crosses, low rates of seed germination, and long crop cycle 2; 3.
On the other hand, genetic engineering techniques are alternative tools for the conventional breeding process, which may overcome some of the issues, including sterility and length for the genetic improvement because several crosses should be performed in conventional breeding2;3. Therefore, the use of breeding technologies, including genetic engineering and recently, genome editing, has revolutionized genetic modification strategies for basic and applied science4. The search for a methodology to obtain in vitro banana transgenic plants in weeks after co-cultivation with Agrobacterium tumefaciens was the objective of this research. Usually, six to twelve months could last to obtain transgenic in vitro banana plants after Agrobacterium-mediated transformation of embryogenic cell suspensions. Furthermore, the development of embryogenic cell suspensions could take eight to twelve months 2; 3; 5; 6; 7. Therefore, the genetic transformation from in vitro banana plants was performed through Agrobacterium tumefaciens; and confirmation of transformation was performed by using the luciferase reporter gene system and PCR.
MATERIALS AND METHODS
Plants preparation
Banana in vitro plants of the cultivar 'Williams' was obtained from the germplasm bank of the Tissue Culture area from CIBE-ESPOL. In vitro, plants were maintained according to Korneva et al. 8.
Agrobacterium strains and plasmids
In this study, the Agrobacterium tumefaciens strain LBA4404 was used, and the binary vectors pLVCIBE1 and pLVCIBE2 were developed containing the gene cassettes PMabHIPP::luc2::Tnos and P35S::luc2::Tnos, respectively 9. The selectable marker gene used was the hpt for hygromycin B resistance, fused to the CaMV 35S promoter.
The plasmids were transferred to agrobacteria by using 1 µg of DNA from plasmid mixed with A. tumefaciens competent cells strain LBA4404. Briefly, the competent cells were frozen together with the plasmid using liquid nitrogen, then a heat shock treatment was given at 37°C for 5 minutes in a water bath, and finally, samples were incubated for 5 minutes on ice. Bacteria were incubated for four hours on YEP liquid medium (peptone 10g/L, yeast extract 10g/L, NaCl 5g/L) at 28ºC on a shaker at 225 rpm. Samples were centrifuged for 30 seconds at 3000g, the supernatant was discarded, and cells were resuspended in 100 µl of YEP liquid medium. The bacterial suspension was cultured into Petri dishes with YEP solid medium with 50 µg/mL of the antibiotics rifampicin (Sigma), and 50µg/mL of kanamycin (Calbiochem). Samples were then incubated at 28ºC for 2 days.
Preparation of Agrobacterium for genetic transformation using 'Williams’ meristematic tissue.
A single colony of A. tumefaciens was inoculated into 5 mL of liquid YEP medium containing kanamycin (50 µg/mL) and rifampicin (50 µg/mL) and grown overnight at 28°C on a rotary shaker at 200 rpm. An aliquot (1 mL) of the overnight culture was inoculated into 50 mL of liquid YEP medium containing the same antibiotics and allowed to grow at 28°C with vigorous shaking until the OD600 reached approximately 0.8 units 10; 11. The bacteria suspension was centrifuged at 5000 g for 10 minutes and resuspended in 25 mL of MS medium without antibiotics for 30 minutes at 25 rpm.
Inoculation and co-cultivation of explants
Co-cultivation of agrobacteria with banana explants was performed in MS culture medium supplemented with acetosyringone (200 μM) without antibiotics for 30 min at 100 rpm. After co-cultivation of agrobacteria with ‘Williams' banana explants, a vacuum infiltration was performed at 400 mmHg for 10 minutes in MS medium supplemented with acetosyringone (200 μM) and maintained for 18 hours at 70 rpm in complete darkness. Two consecutive washes of 60 minutes each in fresh MS liquid medium were performed and maintained in darkness. After washing, the explants were maintained on sterile paper towels for drying, following subculture in semisolid MS medium, and incubation at 21°C for 3 days in complete darkness. After this time, the explants were subcultured in fresh semisolid MS medium, supplemented with Timentin (200 μg/mL) and hygromycin B (12.5 μg/mL) for 15 days at 25°C.
The same co-cultivation methodology was performed using two types of explants; i) whole in vitro plants (approximately 3-cm in height) without any excision, and ii) a bisected 2-cm region cut from the meristematic tissue of in vitro plants. For both cases, the banana cultivar 'Williams' (genotype AAA) was used (Fig. 1).
Luciferase (LUC) activity assay
For the transformation of the whole in vitro banana plants, luciferase activity assays were performed on explants after 15 days under a selection medium containing 200 μg/mL of Timentin and 12.5 μg/mL of hygromycin B (15 days after Agrobacterium infection). In vitro banana plants were transferred to sterilized 9-cm Petri dishes before the application of luciferin. The equipment STELLA3200 (Raytest, Germany) was used to detect LUC activity in complete darkness, using a CCD camera, after 20 µl of luciferin (500 µM) on the banana plants. The acquisition time was 1 min, and the data obtained were analyzed with the AIDA Raytest software (Raytest, Germany).
Selection and regeneration
For the excised bisected meristematic region, selection and regeneration were performed by subculturing on fresh BN semisolid medium supplemented with Timentin (200μg/mL) and hygromycin B (12.5μg/mL), with subculturing every two weeks on fresh medium at 25°C with a 16-h photoperiod for a total of eight weeks.
Polymerase chain reaction to detect transgenic events
To confirm the genetic transformation of the plants regenerated from the excised meristematic region after two months of co-cultivation with A. tumefaciens, a PCR was performed with specific primers to detect the nopaline synthase terminator (TnosF-1: GAATCCTGTTGCCGGTCTTG; and, TnosR-2: TTATCCTAGTTTGCGCGCTA), the expected amplicon size was 180 bp; and the hpt gene (HPTF-1: GATGTTGGCGACCTCGTATT; and, HPTR-2: CACTGGCAAACTGTGATGGA), the expected amplicon size was 216 bp. DNA extraction was performed from leaf tissues using a CTAB modified protocol, according to Pacheco Coello et al. 12. PCR conditions include initial denaturation of 95°C for 3 minutes; then 30 cycles of 95°C for 30 seconds, 57°C for 30 seconds, and 72°C for 30 seconds; with a final extension of 72°C for 5 minutes. The results were visualized on 1.2% agarose gels in 1X TAE buffer.
RESULTS AND DISCUSSION
Once the transformation and subculture process was completed, about 20 shoots were obtained from the meristematic tissue, and they were kept in MS culture medium in Petri dishes following subculture to glass flasks with the same medium changes to fresh culture medium every two weeks. After two weeks of Agrobacterium infection of in vitro plants, luciferase activity was detected in the banana plants transformed, showing expression in all banana tissues (Fig. 2), and seven out of ten putative transgenic lines tested showed PCR amplification (Fig. 3). Both results confirmed the genetic transformation using in vitro banana plants.
The protocol developed will facilitate plants' genetic transformation of different banana and plantain cultivars and other plant species 13. It should also be emphasized that the methodology using Agrobacterium tumefaciens is one of the most used due to its simplicity in terms of techniques and can be used in different types of plant tissues such as those demonstrated in this research 14.
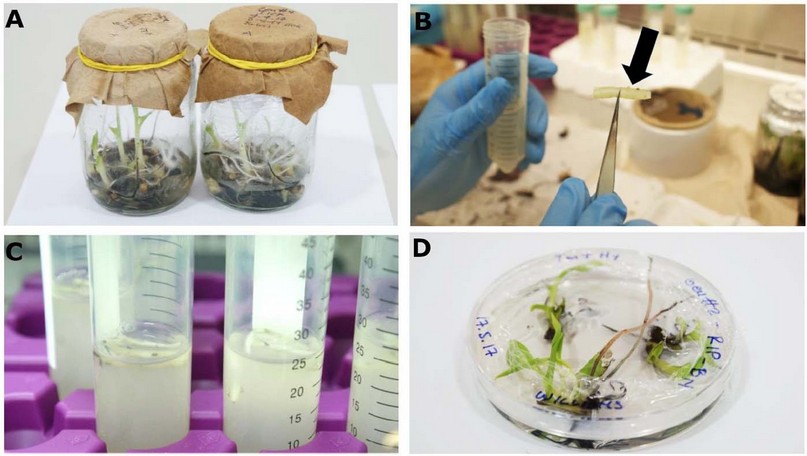

Figure. 1. A. In vitro banana plants from the 'Williams’ cultivar used for transformation. B. Excised bisected banana meristematic tissue region with approximately 2-cm in length (arrow). C. Co-culture process in MS medium supplemented with acetosyringone (200 μM) and Agrobacterium tumefaciens with and O.D.600nm of 0.8 units D. Putative transgenic in vitro plants two months after the meristematic-transformation process in culture medium BN 15.
Although the genetic transformation of embryogenic cell suspension in banana is preferred because transgenic chimeric plants are avoided; several studies developed transformation using meristematic or in vitro plants in Musa varieties (reviewed by López et al.16), indicating that this methodology is useful, which was confirmed in this study using luciferase reporter genes and PCR.
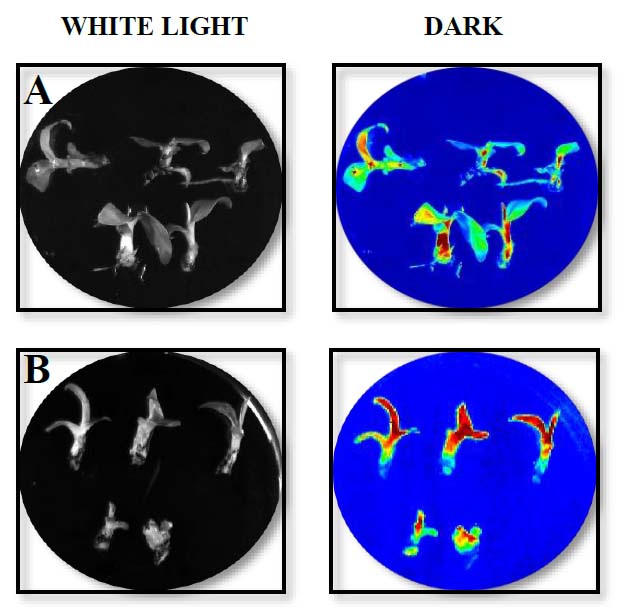
Figure. 2. LUC activity assay of in vitro ‘Williams’ plants after 15 days of co-cultivation with Agrobacterium-mediated transformation of the whole in vitro plants. In vitro banana plants were transformed with vectors pLVCIBE1 [MabHIPP promoter, 9] (A) and pLVCIBE2 (P35S promoter) (B). WHITE LIGHT indicates that the picture was acquired under light conditions, while DARK indicates the picture was acquired under complete darkness.
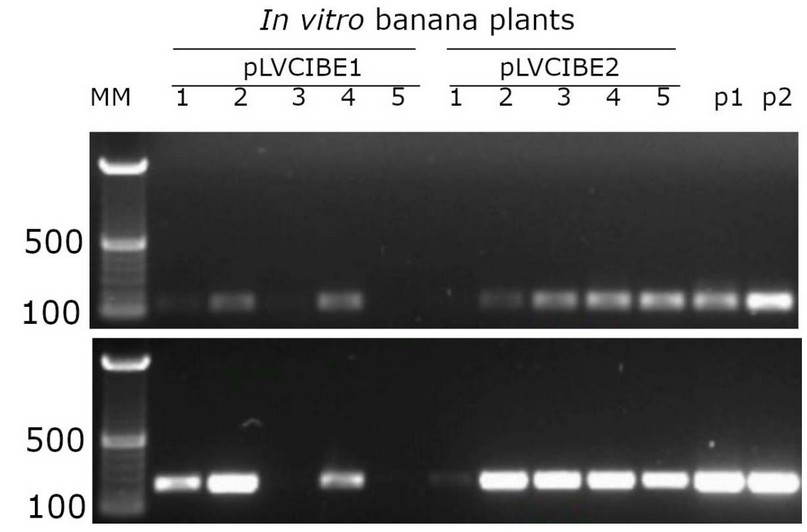
Figure. 3. PCR detection of transgenic banana lines after two months of co-cultivation of A. tumefaciens with the bisected banana in vitro meristems. Agarose gel electrophoresis of PCR using primers to detect the terminator from nopaline synthase (upper level) and the hygromycin-B-phosphotransferase gene (lower level); both sequences contained in the vectors pLVCIBE1 and pLVCIBE2. Numbers at the left indicate reference sizes of DNA in base pairs (500 bp and 100 bp). Numbers 1-5 indicate independent transgenic banana lines for each construct. Vectors DNA for pLVCIBE1 and pLVCIBE2 are indicated as p1 and p2, respectively. MM refers to 100 bp molecular weight markers.
CONCLUSIONS
In this study, we demonstrated the genetic transformation of apical meristems from the Musa cultivar ‘Williams’, allowing the regeneration of transgenic plants in a short period. The Agrobacterium strain LBA4404 could genetically transformed in vitro plants of the banana cultivar ‘Williams’, by using hygromycin B (12.5μg/mL) in the selection process. The luciferase reporter gene system was functional in banana by using the codon-optimized luc2. This transformation methodology could be used for gene functional studies or genetic improvement for (a)biotic stress resistance in banana and plantain.
ACKNOWLEDGMENT
The MabHIPP promoter was cloned from the plasmid pESKUL7 donated by Professor Rony Swennen from KU Leuven.
FUNDING
Genetic transformation experiments were performed in the framework of the Secretaría Nacional de Educación Superior, Ciencia, Tecnología e Innovación (grants PIC-08-0000300 and SENACYT-FWO-10-005); and the VLIR-UOS (grant “Folate Biofortification of Banana Musa spp.” ZEIN2014RIP25).
COMPETING INTEREST
The authors declare no competing interest
REFERENCES
1. Nelson, S. C., Ploetz, R. C., & Kepler, A. K. (2006). Musa species (bananas and plantains). In C. R. Elevitch (Ed.), Species profiles for Pacific Island agroforestry (pp. 1–33). Permanent Agriculture Resources (PAR).
2. Sánchez Timm, L. E., & Santos Ordoñez, E. G. (2011). Estandarización del protocolo de transformación genética de células embriogénicas de banano de la variedad ‘Williams’ (AAA) mediada por Agrobacterium tumefaciens. Revista Tecnológica - ESPOL, 23(1).
3. Santos, E., Sánchez, E., Hidalgo, L., Chávez, T., Villao, L., Pacheco, R., Flores, J., Korneva, S. and Navarrete, O. (2016). Advances in banana transformation through Agrobacterium tumefaciens in Ecuador: progress, challenges and perspectives. Acta Hortic. 1114, 197-202 DOI: 10.17660/ActaHortic.2016.1114.27
4. Altpeter, F., Springer, N. M., Bartley, L. E., Et. al., (2016). Advancing Crop Transformation in the Era of Genome Editing. The Plant cell, 28(7), 1510–1520. https://doi.org/10.1105/tpc.16.00196.
5. Sowmya, H. D., Usharani, T. R., & Mohandas, S. (2016). Somatic embryogenesis as a tool in genetic transformation. In Banana: Genomics and Transgenic Approaches for Genetic Improvement (pp. 141–155). Springer Singapore. https://doi.org/10.1007/978-981-10-1585-4_10
6. Santos, E., Sánchez, E., Hidalgo, L., Chávez, T., Villao, L., Pacheco, R., & Navarrete, O. (2016). Status and challenges of genetically modified crops and food in Ecuador. In Acta Horticulturae (Vol. 1110, pp. 229–235). International Society for Horticultural Science. https://doi.org/10.17660/ActaHortic.2016.1110.33
7. Santos, E., Pacheco, R., Villao, L., Galarza, L., Ochoa, D., Jordán, C., & Flores, J. (2016). Promoter analysis in banana. In Banana: Genomics and Transgenic Approaches for Genetic Improvement (pp. 157–179). Springer Singapore. https://doi.org/10.1007/978-981-10-1585-4_11
8. Korneva, S., Flores, J., Santos, E., Pinã, F., & Mendoza, J. (2013). Plant regeneration of plantain “Barraganete” from somatic embryos using a temporary immersion system. Biotecnologia Aplicada, 30(4), 267–270.
9. Villao, L., Sánchez, E., Romero, C., Galarza, L., Flores, J., & Santos-Ordóñez, E. (2019). Activity characterization of the plantain promoter from the heavy metal-associated isoprenylated plant gene (MabHIPP) using the luciferase reporter gene. Plant Gene, 19. https://doi.org/10.1016/j.plgene.2019.100187
10. Rustagi, A., Jain, S., Kumar, D., Shekhar, S., Jain, M., Bhat, V., & Sarin, N. B. (2014). High Efficiency Transformation of Banana [Musa acuminata L. cv. Matti (AA)] for Enhanced Tolerance to Salt and Drought Stress Through Overexpression of a Peanut Salinity-Induced Pathogenesis-Related Class 10 Protein. Molecular Biotechnology, 57(1), 27–35. https://doi.org/10.1007/s12033-014-9798-1.
11. Tripathi, L., Tripathi, J.N. & Hughes, J.D. (2005). Agrobacterium-mediated transformation of plantain (Musa spp.) cultivar Agbagba. African Journal of Biotechnology, 4(12), 1378-1383.
12. Pacheco Coello, R., Pestana Justo, J., Factos Mendoza, A. et al. Comparison of three DNA extraction methods for the detection and quantification of GMO in Ecuadorian manufactured food. BMC Res Notes 10, 758 (2017). https://doi.org/10.1186/s13104-017-3083-x.
13. Hwang, H.-H., Yu, M., & Lai, E.-M. (2017). Agrobacterium -Mediated Plant Transformation: Biology and Applications. The Arabidopsis Book, 15, e0186. https://doi.org/10.1199/tab.0186.
14. Díaz Granados, Cristina, & Chaparro-Giraldo, Alejandro. (2012). MÉTODOS DE TRANSFORMACIÓN GENÉTICA DE PLANTAS. Revista U.D.C.A Actualidad & Divulgación Científica, 15(1), 49-61. Retrieved September 22, 2020, from http://www.scielo.org.co/scielo.php?script=sci_arttext&pid=S0123 42262012000100007&lng=en&tlng=es.
15. Ortega Pérez, N. V., Korneva Korneva, S., Santos Ordóñez, E. G., Ruíz Barzola, O., & Peralta García, E. L. (2011). Obtención de Multimeristemos y Callos de Diferentes Variedades de Banano y Plátano (Musa spp.) a partir de “Meristemos Apicales” y “Scalps”. Revista Tecnológica - ESPOL, 23(1).
16. López J., Santos-Ordoñez E., González L. (2020) Complementation of Bananas Conventional Breeding Programs Through Biotechnological Genetic Improvement. In: Chong P., Newman D., Steinmacher D. (eds) Agricultural, Forestry, and Bioindustry Biotechnology and Biodiscovery. Springer, Cham. https://doi.org/10.1007/978-3-030-51358-0_3
Received:5 October
Accepted: 10 November 2020
Liliana Villao1, José Flores1, 2, Efrén Santos-Ordóñez 1, 3
1ESPOL Polytechnic University, Escuela Superior Politécnica del Litoral, ESPOL, Centro de Investigaciones Biotecnológicas del Ecuador (CIBE), Campus Gustavo Galindo Km 30.5 Vía Perimetral, P.O. Box 09-01-5863, Guayaquil, Ecuador. ORCID: orcid.org/0000-0002-2677-1415
2Facultad de Ciencias Naturales, Universidad de Guayaquil, Av. Raúl Gómez Lince s/n Av. Juan Tanca Marengo, Ecuador. ORCID: orcid.org/0000-0003-2909-7849
3ESPOL Polytechnic University, Escuela Superior Politécnica del Litoral, ESPOL, Facultad de Ciencias de la Vida (FCV), Campus Gustavo Galindo Km 30.5 Vía Perimetral, P.O. Box 09-01-5863, Guayaquil, Ecuador. ORCID: orcid.org/0000-0002-0749-6024
Corresponding author: [email protected]
